Thousands of species are polyploid. However, the long-term establishment of organisms that have undergone ancient whole genome duplications (WGDs) has been exceedingly rare and when we analyse the genomes of plants and animals, we can, at most, find evidence for a very limited number of WGDs that survived on the longer term. The paucity of (established) ancient genome duplications and the existence of so many species that are currently polyploid provides a fascinating paradox. There is growing evidence that the majority of ancient WGDs were established at specific times in evolution, for instance during periods of environmental change and periods of mass-extinction. The reason for this ‘stress’-polyploidy relationship has been the subject of considerable speculation and several hypotheses have been put forward to explain this observation: (a) stressful conditions promote polyploid formation; (b) polyploidisation causes a niche shift allowing polyploids to grow in conditions that are unsuitable for their non-polyploid ancestors; and (c) polyploids have an increased evolvability and consequently adapt faster to a changing environment. In future work, we wants to unravel the mechanistic underpinnings of why and how polyploids can outcompete non-polyploids. We will address these questions by replaying the ‘genome duplication tape of life’ in two different model systems, namely Chlamydomonas and Spirodela (duckweed). We will run long-term evolutionary (and resequencing) experiments that will be complemented with in-silico experiments based on so-called digital organisms (DOs) running on artificial genomes. Complementary modelling approaches will also be employed to study the effects of polyploidy from an eco-evolutionary dynamics perspective. By integrating the results obtained from these in vivo and in silico experiments, we hope to obtain important novel insights in the adaptive potential of polyploids under stressful conditions or during times of environmental and/or climate change.
Selected Publications:
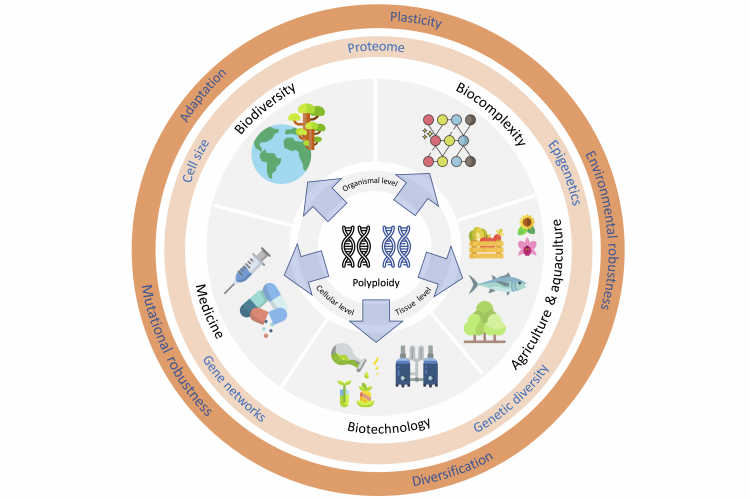
Fox, Donald T., Douglas E. Soltis, Pamela S. Soltis, Tia-Lynn Ashman, and Yves Van de Peer. 2020. “Polyploidy: A Biological Force From Cells to Ecosystems.” Trends in Cell Biology, July, S0962892420301240. https://doi.org/10.1016/j.tcb.2020.06.006.
Zwaenepoel, Arthur, and Yves Van de Peer. 2019. “Inference of Ancient Whole-Genome Duplications and the Evolution of Gene Duplication and Loss Rates.” Edited by Stephen Wright. Molecular Biology and Evolution 36 (7): 1384–1404. https://doi.org/10.1093/molbev/msz088.
Zwaenepoel, Arthur, and Yves Van de Peer. 2019. “Wgd—Simple Command Line Tools for the Analysis of Ancient Whole-Genome Duplications.” Edited by John Hancock. Bioinformatics 35 (12): 2153–55. https://doi.org/10.1093/bioinformatics/bty915.
Van de Peer, Yves, Eshchar Mizrachi, and Kathleen Marchal. 2017. “The Evolutionary Significance of Polyploidy.” Nature Reviews Genetics 18 (7): 411–24. https://doi.org/10.1038/nrg.2017.26.
Lohaus, Rolf, and Yves Van de Peer. 2016. “Of Dups and Dinos: Evolution at the K/Pg Boundary.” Current Opinion in Plant Biology 30 (April): 62–69. https://doi.org/10.1016/j.pbi.2016.01.006.
Vanneste, Kevin, Guy Baele, Steven Maere, and Yves Van de Peer. 2014. “Analysis of 41 Plant Genomes Supports a Wave of Successful Genome Duplications in Association with the Cretaceous–Paleogene Boundary.” Genome Research 24 (8): 1334–47. https://doi.org/10.1101/gr.168997.113.
Vanneste, Kevin, Steven Maere, and Yves Van de Peer. 2014. “Tangled up in Two: A Burst of Genome Duplications at the End of the Cretaceous and the Consequences for Plant Evolution.” Philosophical Transactions of the Royal Society B: Biological Sciences 369 (1648): 20130353. https://doi.org/10.1098/rstb.2013.0353.